Data Fields
The Following list shows the data that is available to approved research projects in addition to the Epidemiology Questionnaire
| Table | Field | Description | Available values |
|---|---|---|---|
| BloodSpecimen | DateTaken | Date the blood was drawn from the participant | |
| BloodSpecimen | NonBloodType | Sample type collected if not blood (usually mouthwash) | |
| BloodSpecimen | CurrentlyPregnant | Was the participant pregnant at the time the blood was taken | |
| BloodSpecimen | DateLastMenstrual | Date of last menstrual period when blood was taken | |
| BloodSpecimen | AveDaysMenstrualCycle | Average number of days of menstrual cycle for participant | |
| BloodSpecimen | Menopause | Had the participant reached menopause at the time the blood was taken | |
| BloodSpecimen | OCP | Was the participant taking the oral contraceptive pill at the time the blood was taken | |
| Cancer | CancerSite | Site of cancer | |
| Cancer | Laterality | Laterality of cancer | |
| Cancer | Diagnosed | Date diagnosed | |
| Cancer | DiagnosedApprox | Was diagnosis date approximate | |
| Cancer | DiagnosisDateUnknown | Diagnosis date not able to determined (Diagnosed will not be given if this field is true) | |
| Cancer | VerificationOutcome | Was the cancer able to be verified | |
| Death | CancerCause | Did the participant die from cancer | |
| Death | CancerSite | Site of the cancer that contributed to the participant's death | |
| Family | UFN | Unique family ID | |
| Family | CriteriaOnEntry | Straddie criteria the family was recruited on | |
| Family | FamilyReviewDate | Date at last family review | |
| Family | CriteriaOnReview | Straddie criteria at last review | |
| FamilyMutation | Nomenclature | Description of classified mutation | |
| FamilyMutation | Gene | Gene of classified mutation | |
| FamilyMutation | MutationEffect | Effect of classified mutation | |
| Haplotype BRCA1 | UFN | Family number | |
| Haplotype BRCA1 | Kindred type | Kindred type | |
| Haplotype BRCA1 | BRCA1 shared/typed | BRCA1 shared/typed | |
| Haplotype BRCA1 | Date | Date tested | |
| Haplotype BRCA1 and BRCA2 | UFN | Family number | |
| Haplotype BRCA1 and BRCA2 | Kindred type | Kindred type | |
| Haplotype BRCA1 and BRCA2 | BRCA1 shared/typed | BRCA1 shared/typed | |
| Haplotype BRCA1 and BRCA2 | Date | Date tested | |
| Haplotype BRCA2 | UFN | Family number | |
| Haplotype BRCA2 | Kindred type | Kindred type | |
| Haplotype BRCA2 | BRCA2 shared/typed | BRCA2 shared/typed | |
| Haplotype BRCA2 | Date | Date tested | |
| Haplotype Neither | UFN | Family number | |
| Haplotype Neither | Kindred type | Kindred type | |
| Haplotype Neither | Date | Date tested | |
| Mutation | Report Date | Date the mutatation was reported | |
| Mutation | Gene | Gene tested | |
| Mutation | MutationResult | Result of the mutation test (positive or negative) | |
| Mutation | ResultType | Diagnostic (clinical) or research (kconfab) mutation test | |
| Mutation | TestMethod | The method used for the test | |
| Mutation | Predictive | Was the test a predictive test | |
| Mutation | FullScreen | Was the whole gene tested | |
| Mutation | Exon13 | Was Exon 13 tested | |
| Mutation | LargeDeletion | Was the test a search for a large deletion | |
| Mutation | MutationDesc | Description of the mutation as given by the testing laboratory | |
| Mutation | FamilyMutation | Link to the Family mutation the test relates to | |
| Pathology | Prophylactic | Was the procedure for prophylactic treatment | |
| Pathology | Side | Side of the body the tissue was taken from | |
| Pathology | Report Date | Date the report was generated by the pathology laboratory | |
| Pathology | PathologyProcedure | Procedure that extracted the sample that was tested | |
| Pathology | PathSampleType | Type of tissue that was extracted | |
| Pathology | Tumour | Did the tested sample contain tumour | |
| Pathology | TumourSize | Size of tumour found | |
| Pathology | Topography | Topography classification (ICD-O description) | |
| Pathology | Morphology | Morphology classification (ICD-O description) | |
| Pathology | Behaviour | Behaviour classification | |
| Pathology | Grade | Grade classification | |
| Pathology | Oestrogen | Oestrogen receptor test result | |
| Pathology | Progesterone | Progesterone receptor test result | |
| Pathology | CA125 | CA125 test result | |
| Pathology | HER2 | HER2 test result | |
| Pathology | PSA | PSA test result | |
| Pathology | Gleason | Gleason score | |
| Pathology | NodesTested | Number of lymph nodes tested | if blank then not detailed in report |
| Pathology | NodesPositive | Number of positive lymph nodes | |
| Pathology | DCISMorphology | Morphology of any in-situ carcinoma found in addition to invasive | |
| Pathology | DCISGrade | Grade of any in-situ carcinoma found in addition to invasive | |
| PathologyReview | ReviewDate | Date of the pathology review | |
| PathologyReview | SourceLab | Pathology laboratory who generated the original pathology report | |
| PathologyReview | TumourSize | Tumour diameter recorded on report (mm) | |
| PathologyReview | HR | Were hormone receptor tested | |
| PathologyReview | HRMethod | Method used to test the hormone receptors | |
| PathologyReview | ER | Oestrogen receptor “Positive” or “Negative” and quantitative result (if supplied) | |
| PathologyReview | PR | Progesterone receptor “Positive” or “Negative” and quantitative result (if supplied) | |
| PathologyReview | LNNumber | Number of lymph nodes identified | |
| PathologyReview | MetNum | Number of lymph nodes containing metastases | |
| PathologyReview | SentinalSelect | Staining results for sentinal nodes | |
| Pathology review has been performed only on 214 slide review samples | |||
| Person | UPN | Unique person ID | |
| Person | Gender | Gender of participant | |
| Person | StartDate | Date the participant's details were added to the database | |
| Person | DOB | Date of birth of the participant | |
| Person | DOBUnknown | Is the date of birth approximate | |
| Person | DeathDate | Date of death of the participant | |
| Person | DeathDateApprox | Is the date of death approximate | |
| Person | NoFamilyHistory | The participant has no family history of their own | |
| PersonOtherStudy | OtherStudy | Description of other study the participant is enrolled in | |
| PersonRelation | ParentID | ID of the participant's parents | |
| Family pedigrees are also available in .jpg or Progeny format | |||
| QANormalBreast | ArchPattern | Silverberg architectural pattern | |
| QANormalBreast | NuclearPleo | Silverberg nuclear pleopmorphism | |
| QANormalBreast | Mitotic | Silverberg mitotic activity | |
| QANormalBreast | Cellularity | Silverberg cellularity | |
| QANormalBreast | Neoplastic | % neoplastic component | |
| QANormalBreast | Lymph | Non-neoplastic component: % lymph | |
| QANormalBreast | Stroma | Non-neoplastic component: % stroma | |
| QANormalBreast | NormalType1Percent | Normal epithelium component: % type 1 | |
| QANormalBreast | NormalType2Percent | Normal epithelium component: % type 2 | |
| QANormalBreast | Necrotic | % necrotic component | |
| QANormalBreast | Morphology | Morphology description | |
| QANormalBreast records QA results for normal (prophylactic) breast tissue. | |||
| QANormalOvary | Side | Tissue laterality | |
| QANormalOvary | SurfaceEpi | Presence of surface epithelium | |
| QANormalOvary | EpiAmount | Amount of surface epithelium | |
| QANormalOvary | Pappilations | Presence of surface epithelium papillations | |
| QANormalOvary | PapAmount | Amount of epithilium papillations | |
| QANormalOvary | Cysts | Presence of epithelial inclusion cysts | |
| QANormalOvary | CystAmount | Amount of epithilial cysts | |
| QANormalOvary | Invaginations | Presence of cortical invaginations | |
| QANormalOvary | InvagAmount | Amount of cortical invaginations | |
| QANormalOvary | Stroma | Presence of stroma | |
| QANormalOvary | Follicles | Presence of cortext follicles | |
| QANormalOvary | CorpLutea | Presence of cortex corpora lutea | |
| QANormalOvary | FollCysts | Presence of follicular cysts | |
| QANormalOvary | Rete | Presence of rete ovarii | |
| QANormalOvary | Fallopian | Presence of fallopian tube | |
| QANormalOvary | Morphology | Preservation of morphology | |
| QANormalOvary | %Tumour | % tumour in sample | |
| QANormalOvary records QA results for normal (prophylactic) ovarian tissue | |||
| QAProstate | Lab | Pathology laboratory from which the slide was sourced | |
| QAProstate | Histology | Prostate histology | |
| QAProstate | SpecimenType | Specimen type | |
| QAProstate | SpecimenBiopsyCores | If specimen type is biopsy; the number of cores | |
| QAProstate | SpecimenBiopsyCoresTumour | If specimen type is biopsy; the number of cores with tumour | |
| QAProstate | TumourVolume | Tumour Volume | |
| QAProstate | GleasonScore1 | Gleason score: primary pattern | |
| QAProstate | GleasonScore2 | Gleason score: secondary pattern | |
| QAProstate | PrimaryTumour | If no tumour | |
| QAProstate | PrimaryTumourT1 | Extent of clinically inapparent tumour | |
| QAProstate | PrimaryTumourT2 | Extent of tumour confined within prostate | |
| QAProstate | PrimaryTumourT3 | Tumour extends through prostate capsule | |
| QAProstate | PrimaryTumourT4 | Tumour is fixed or invades adjacent structures | |
| QAProstate | RegionalNodes | Status of regional lymph nodes | |
| QAProstate | DistantMetastasis | Distant metastasis | |
| QAProstate | ResectionMargins | Involvement of resection margins | |
| QAProstate | Extra-prostatic extension | Extent of extra-prostatic extension | |
| QAProstate | PerineuralInvasion | Presence of perineural invasion | |
| QAProstate | NeurovascularBundles | Involvement of neurovascular bundles | |
| QAProstate | Lymphovascular invasion | Presence of lymphovascular invasion | |
| QAProstate | NeuroendocrineFeatures | Presence of neuroendocrine features | |
| QAProstate | IntraEpithelialNeoplasia | Presence of high grade prostatic intraepithelial neoplasia | |
| QAProstate | OtherFeatures | Any other histological features | |
| QAProstate records QA results for prostate tissue | |||
| QATumour | TissueType | Type of tissue QA'd | |
| QATumour | Grade | Tumour grade | |
| QATumour | ArchPattern | Silverberg architectural pattern (ovary only) | |
| QATumour | NuclearPleo | Silverberg nuclear pleopmorphism (ovary only) | |
| QATumour | Mitotic | Silverberg mitotic activity (ovary only) | |
| QATumour | Neoplastic | % neoplastic component | |
| QATumour | Lymph | Non-neoplastic component: % lymph | |
| QATumour | Stroma | Non-neoplastic component: % stroma | |
| QATumour | NormalType1Percent | Normal component: % type 1 | |
| QATumour | NormalType2Percent | Normal component: % type 2 | |
| QATumour | Necrotic | % necrotic component | |
| QATumour | Morphology | Morphology description | |
| QATumour | BRCAXReview | BRCA-like features (breast only) | |
| QATumour records QA results for tumour tissue (not just breast) | |||
| SlideReview | SourceLab | Pathology laboratory from which the slide was sourced | |
| SlideReview | AccessionNumber | Laboratory accession number | |
| SlideReview | SlidesExamined | Which slides were examined | |
| SlideReview | ObsDate | Review date | |
| SlideReview | HistoSubType | Histological subtype of invasive carcinoma | |
| SlideReview | LobType | Lobular type | |
| SlideReview | Tubule | Extent of tubule formation | |
| SlideReview | NuclearPleo | Extent of nuclear pleomorphism | |
| SlideReview | Mitotic | Total mitotic figures in 10 high power fields | |
| SlideReview | MitoticTotal | Mitotic score (1, 2 or 3) | |
| SlideReview | Grade | Overall grade | |
| SlideReview | SolidSheet | % of tumour as solid sheet of cells | |
| SlideReview | Circumference | % circumference continuous pushing margin | |
| SlideReview | Necrosis | Confluent necrosis (“Present” or “Absent”) | |
| SlideReview | Infiltrate | Tumour lymphocytic infiltrate | |
| SlideReview | Borders | Tumour cells: presence of discernable cell borders | |
| SlideReview | Vesicular | Tumour cells: presence of vesicular nuclei | |
| SlideReview | EoNucleoli | Tumour cells: presence of prominent eosinophilic nucleoli | |
| SlideReview | DCISMass | % DCIS | |
| SlideReview | DCISExtending | Is the DCIS extending beyond the confines of the main tumour mass | |
| SlideReview | DCISGrade | DCIS nuclear grade | |
| SlideReview | DCISNecrosis | DCIS necrosis | |
| SlideReview | DCISPolarisation | Presence of DCIS cell polarisation | |
| SlideReview | DCISArchitecture | DCIS architectural pattern (multiple selections) | |
| SlideReview | LCIS | Presence of LCIS | |
| SlideReview | BRCA1 | Is the sample consistent with a BRCA1 phenotype | |
| Slide review has been performed by a pathologist on 214 slides from BRCAX families | |||
| TestSensitivity | Gene | Gene tested | |
| TestSensitivity | Sensitivity | Sensitivity of all mutation testing performed on participant | |
| See Senstivitities for a description of each test sensitivity category | |||
| TissueAliquot | TissueSize | The aliquot size | |
| TissueAliquot | SampleType | Type of sample stored | |
| TissueAliquot | Side | Side of the body the tissue was taken from | |
| TissueAliquot | Tumour | Is there tumour present in the aliquot | |
| TissueCollection | TissueType | The type of tissue collected | |
| TissueCollection | DateTaken | The date the tissue was collected | |
| TissueCollection | Prophylactic | Was the tissue removed for prophylactic reasons | |
| TissueCollection | TimeToFrozen | The time taken to freeze the tissue | |
| TissueCollection records details of a surgical collection event. Each collection may be divided into a number of aliquots which are recorded in the TissueAliquot table | |||
| Treatment | Treatment | Description of non-cancer-related procedure | |
| Treatment | Reason | Was the treatment cancer-related or prophylactic. | |
| Treatment | StartDate | Date the treatment began | |
| Treatment | EndDate | Date the treatment finished | |
| Treatment | Drug | Treatment drug | |
| Treatment | Dose | Treatment dose | |
Note that specific data can only be provided for participants who were deceased at time of family enrollment, or who have signed a consent form
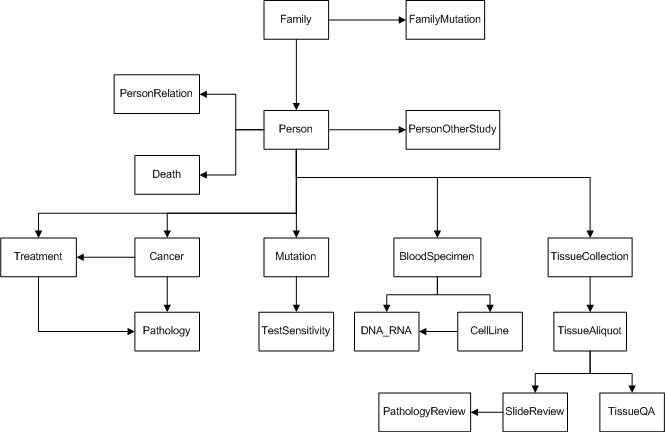
The following diagram illustrates the relationship between tables